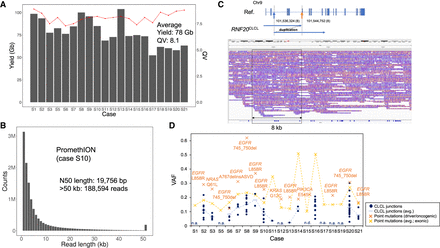
Analysis of lung cancer clinical samples. (A) Sequencing yields and mean quality values of the data sets from PromethION. (B) Raw read length distribution of PromethION sequencing in case S10 (two flow cells; 28×). The N50 length and the number of reads >50 kb are shown in the inset. (C) The CLCL of RNF20 in case S8. An aberrant structure of RNF20 is shown in the upper panel. The structure was a tandem duplication between the junctions (blue arrow and yellow arrow). IGV visualization is represented in the lower panel. The aberrant structure (∼8 kb) is indicated in the box. (D) Variant allele frequencies of CLCL junctions of 20 clinical samples. Variant allele frequencies (VAFs) of CLCL junctions (blue) are shown in comparison with those of point mutations (yellow). VAFs were calculated using Illumina short-read data by Genomon and GenomonSV (VAF > 0.01). Of the 95 CLCL junctions, 79 junctions were also detected in GenomonSV. Driver/oncogenic mutations (known mutations in EGFR, KRAS, NRAS, and PIK3CA) were detected in 13 cases, and their VAFs are presented in orange. No CLCLs were detected in six cases (S1, S5, S12, S17, S18, and S19). (n.a.) Not available; (n.d.) not detected by GenomonSV; thus, the VAF should be considerably low. The estimated VAFs of SNVs and SVs are simultaneously plotted on the same planner for the purpose of comparison, although the accuracy for the VAFs of SVs is not fully validated.











