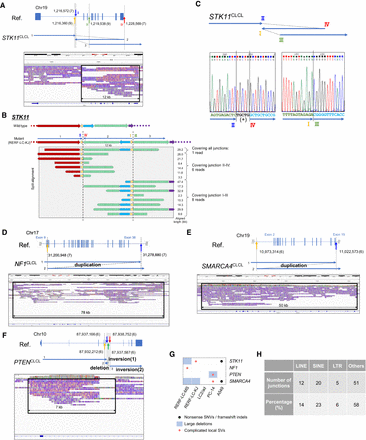
Identification and characterization of complicated local structural aberrations. (A) Structure of STK11 in RERF-LC-KJ. The STK11 CLCL was constructed by a combination of local inversions. We can trace the structure following the ordered arrows, and the junctions are indicated by colored arrows. The SV spanned 12 kb in the human reference genome. (B) Information on long reads for the complicated structure of the STK11 CLCL. The SV structure and aligned reads of PromethION are shown in the upper and lower panels. Aligned length is represented in the margin. (C) The results of Sanger sequencing of junctions of STK11 CLCL. The CLCL junctions of STK11 in RERF-LC-KJ were validated by Sanger sequencing. The CLCL structure is shown in the top panel. The 5-bp insertion was observed in junction I/III. PCR and sequencing primers are shown in the Methods section. (D) Structure of the NF1 in RERF-LC-MS. The structure of the SV was a tandem duplication between the junctions (indicated by a yellow arrow and a blue arrow). The SV spanned 78 kb in the reference genome. (E) Structure of the SMARCA4 in PC-14. The structure of the SV was a tandem duplication of the junctions (indicated by a yellow arrow and a blue arrow). The SV spanned 50 kb in the reference genome. (F) Aberrant genomic structure of PTEN in PC-14. The structure of the SV was a combination of a local inversion and deletion. We can trace the SV structure following the ordered arrows, and the junctions are indicated by colored arrows. The SV spanned 7 kb in the reference genome. (G) Summary of mutation types of four cancer-related genes in five cell lines. (H) The number of complicated SV junctions in each category of genomic context.











