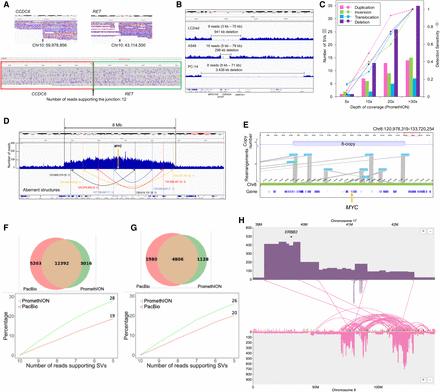
Structural variant analysis of cancer genomes. (A) IGV image of the CCDC6-RET fusion gene of LC2/ad MinION sequencing. The CCDC6-RET fusion gene is a driver mutation of LC2/ad cells. The number of reads supporting the junction is 12. (B) IGV image of a large deletion including CDKN2A of LC2/ad, A549, and PC-14. The deletion of LC2/ad spanned ∼941 kb. The deletion of A549 spanned ∼296 kb. The deletion of PC-14 spanned ∼3438 kb. The parentheses indicate the range of length of reads supporting the deletions. (C) The number of SVs and sensitivities of SV detection in each depth of coverage of PromethION reads. We randomly selected reads to collectively sum up to the indicated sequencing coverage. (D) Depth plotting around the MYC gene of LC2/ad. Arrows indicate junction candidates of the amplification supported by MinION and Illumina reads. Parentheses indicate the number of supporting MinION reads (left) and Illumina paired-end reads (right). (E) The eight-copy MYC region of LC2/ad represented by the optical mapping method. Optical maps with rearrangements and Chromosome 8 (reference) are represented in light blue and yellow-green, respectively. The orange arrow indicates the MYC gene. (F) Comparison of SVs of SK-BR-3 cells from PacBio data (red; previous report) with PromethION data (green). A total of 12,392 SVs were common between the two data sets (upper panel). Percentage of unique SVs overlapping with the other data set when the number of required reads supporting SVs decreased. Colors of the lines correspond to the colors of the regions in the upper Venn diagram (lower panel). (G) Comparison of genic SVs of SK-BR-3 cells from PacBio data with PromethION data in the same manner as F. (H) Large-scale translocations from the region around the ERBB2 gene to Chromosome 8 detected by the PromethION data. The translocations corresponded to the previous report using PacBio data.











