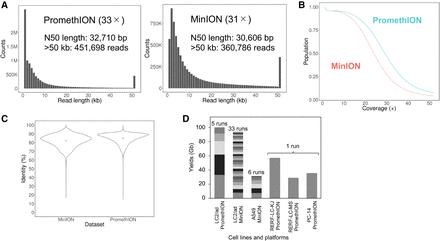
Long-read sequencing of cancer genomes. (A, left) Raw read length distribution of LC/2ad PromethION sequencing. (Right) Raw read length distribution of LC2/ad MinION sequencing. Both PromethION and MinION data sets have many long reads (e.g., >50 kb, 452 kilo reads, and 361 kilo reads, respectively). (B) Cumulative depth curve of LC2/ad. Blue indicates PromethION; red, MinION. More than 50% of the human genome was covered by a sequencing depth of >20× in both data sets. (C) Violin plots of sequence identity of LC2/ad MinION sequencing (left) and PromethION sequencing (right). The points in the violin plots indicate the mean identities of the MinION and PromethION data (82.1% and 84.8%, respectively) (Table 1). The identities were concentrated at >80% in both data sets. (D) Sequencing yields for each data set of PromethION and MinION. Yields from individual flow cells are represented by different colors.











