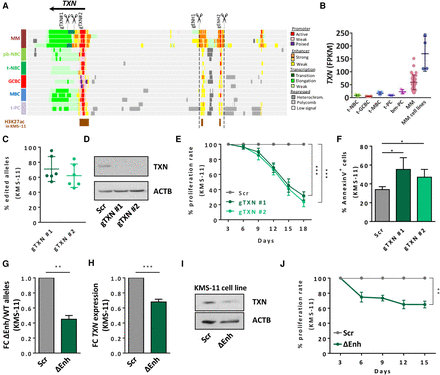
TXN de novo activated enhancer as an essential regulatory element in MM. (A) Genome browser snapshot of the TXN locus and the associated enhancer de novo activated in MM located at 50 kb from the promoter region. Displayed tracks represent the chromatin state annotation in MM patients and normal B cells and an additional track of H3K27ac peaks in KMS-11 cell line (ENCODE Consortium, ENCSR715JBO). gRNA design strategy for TXN knockout and enhancer deletion is also shown. (B) TXN expression in MM patients and cell lines analyzed by RNA-seq. (C) Estimation of the allelic cell population percentage exhibiting indel events in the targeted site, analyzed by Tracking of Indels by DEcomposition (TIDE) web tool. (D) Validation of TXN reduced expression by western blot analysis in KMS-11 cell line. (E) Cell proliferation assay comparing the growth proliferation rates of scramble cells (Scr) and cells harboring two different gRNAs, as determined by flow cytometry analysis. (F) Effect of TXN reduced expression in cell apoptosis, as determined by Annexin V flow cytometry analysis. (G) Quantification of TXN enhancer deletion by genomic DNA qPCR normalized to a distal nontargeted genomic region, represented as fold change of deleted enhancer (ΔEnh) versus wild-type (WT) alleles. (H) TXN mRNA expression levels determined by RT-qPCR. (I) TXN protein levels in cells with deleted enhancer region (ΔEnh) and scrambled cells (Scr) determined by western blot. (J) Cell proliferation assay comparing the growth proliferation rates of scramble cells and cells harboring the enhancer deletion (ΔEnh), as determined by flow cytometry analysis. (*) P < 0.05, (**) P < 0.01, (***) P < 0.001.











