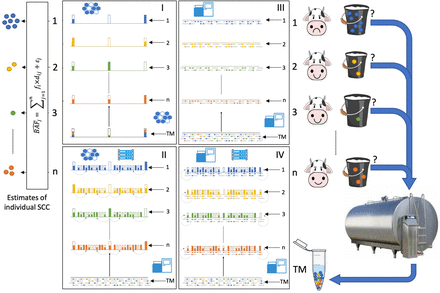
Estimating somatic cell counts (SCCs) in the milk of individual cows by analyzing a sample of milk from the farm's tank. Cows
1 to n contribute different amounts of milk (buckets of various sizes in the figure) to the farm's tank. The milk contains somatic
cells (shown as small spheres in the milk colored by cow) whose numbers reflect the health status of the cow's udder. Cow
1 has higher a SCC, an indicator of subclinical mastitis. SCCs are unknown upon milking (indicated by a question mark). Cows
are individually genotyped once. In scheme I, this is performed using SNP arrays (illustrated by the mesh), yielding genotype
information for the limited number of interrogated SNPs (high bars) that can be summarized by the B-allele frequency as shown (white: 0; half-colored: 0.5; fully colored: 1). SNP genotypes of individual cows are coded in
the same colors as the SCCs. In scheme II, the genotypes of the interrogated SNPs are augmented by imputation (illustrated
by the computer rack), yielding dosage information (B-allele frequency) for many more SNPs (small bars). In scheme III, cows are genotyped individually by shallow whole-genome
sequencing (SWGS; illustrated by the sequencer). Sequence reads (gray lines) are aligned to the reference genome, and alternate
alleles at SNP positions are highlighted as color-coded tics. The B-allele frequency at specific SNP positions is measured as the ratio of the number of reads with the B allele versus the total number of reads. In scheme IV, the genotype information from SWGS is augmented by imputation improving
the accuracy of the B-allele frequency estimates for millions of SNPs (small bars). A small sample of milk (tank milk [TM]) is periodically (e.g.,
monthly or weekly) collected from the farm's tank. DNA is extracted from the TM and genotyped using SNP arrays (scheme I)
or SWGS (schemes II, III, and IV). B-allele frequency for SNP j in the milk (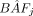 ) is estimated from the ratio of fluorescence intensities when using SNP arrays or from the proportion of reads with B allele in SWGS. The SCCs of individual cows are estimated from a set of linear equations modeling
) is estimated from the ratio of fluorescence intensities when using SNP arrays or from the proportion of reads with B allele in SWGS. The SCCs of individual cows are estimated from a set of linear equations modeling 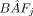 as the sum of B-allele dosage (dij) multiplied by the proportion of the DNA in the tank contributed by cow i (fi). The estimated proportions of DNA contributed by each cow correspond to the values of fi's that minimize the sum of squared errors (ɛj) over all SNPs. The SCCs for individual cows, per se, can be estimated as SCCi = SCCtank × Vtank × fi/Vi, where SCCtank is the SCC measured in the farm's tank, and Vi/Vtank is the proportion of the milk volume contributed by cow i.
as the sum of B-allele dosage (dij) multiplied by the proportion of the DNA in the tank contributed by cow i (fi). The estimated proportions of DNA contributed by each cow correspond to the values of fi's that minimize the sum of squared errors (ɛj) over all SNPs. The SCCs for individual cows, per se, can be estimated as SCCi = SCCtank × Vtank × fi/Vi, where SCCtank is the SCC measured in the farm's tank, and Vi/Vtank is the proportion of the milk volume contributed by cow i.











