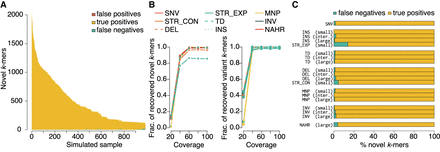
Simulation-based evaluation of novel k-mer detection and subsequent reassembly quality for contigs spanning novel k-mers in error-containing short-read data. (A) Number of k-mers in the progeny correctly identified as novel (true positives), undetected (false negatives), and misidentified as novel (false positives). (B) Novel and variant k-mer recovery for all in silico progeny at simulated mean coverages of 20×, 40×, 60×, 80×, and 100×. (C) For all simulated alleles, the fraction assembled completely (i.e., wholly contained within a single contig) and incompletely (i.e., only partially reconstructed).











