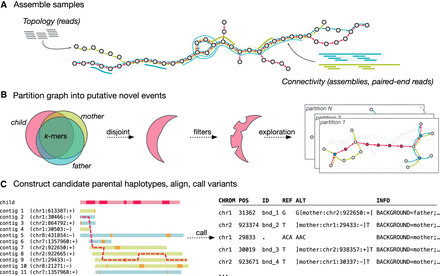
Overview of the Corticall algorithm. (A) Samples are assembled into a multicolor linked de Bruijn graph (LdBG). Short, accurate reads are used to determine graph topology. Longer sequences derived from paired-end reads or from draft/finished assemblies are thread through the graph, providing information on connectivity to overcome repeats but not adding novel k-mers. (B) Novel k-mers, sequences present in the progeny and absent in the parents, are filtered and then used to signal the presence of putative de novo mutations (DNMs). Subgraphs around such events are extracted, forming a set of variant candidates. (C) Regions flanking novel k-mers are assembled to reveal candidate parental haplotypes. The progeny's contig is probabilistically aligned to the set of candidate parental contigs, allowing for mismatches, indels, and (potentially nonallelic) recombination. The resulting alignment thus specifies parental background and (if reference sequences are available) coordinate information. Variants (SNVs, MNVs, indels, translocation breakends, etc.) within the novel k-mer regions are returned as likely DNMs.











