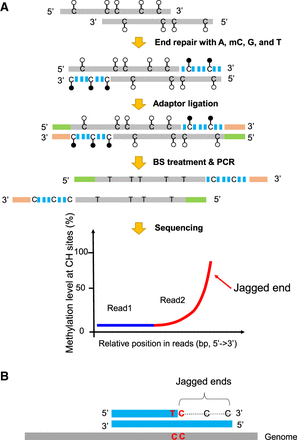
Schematic illustration of plasma DNA jagged end detection using methylation levels at CH sites. (A) The general principle for detecting the presence of jagged ends. In the process of end repair, a DNA molecule carrying 5′ protruding ends (i.e., jagged ends) would be filled up by A, T, mC (methylated C), G with the Klenow fragment without 3′ → 5′ exonuclease activity. The CH (H: A, C, or T) sites present in the newly generated strand were expected to be methylated, whereas the CH sites in the original sequence of the same molecule were generally unmethylated. After the end repair of plasma DNA, the resultant blunted molecules were subjected to bisulfite treatment and PCR amplification. If a molecule contained jagged ends with a 5′ protruding single strand, the methylation levels at CH sites proximal to the 3′ ends (e.g., read2) would be higher than that close to the 5′ end (e.g., read1). Filled lollipops represent methylated Cs, and unfilled lollipops represent unmethylated Cs. The dashed blue lines represent newly filled-up nucleotides. (B) The exact jagged end length deduction. A molecule containing two consecutive cytosines (i.e., “CC” tag) would provide a possibility to directly determine the exact length of a jagged end, when the below criteria are fulfilled: The first one is located within the double-stranded DNA and the other just corresponds to the first nucleotide of the jagged end. Such a “CC” tag would be converted into a “TC” tag after the bisulfite conversion followed by PCR amplification. Therefore, the dashed line indicates the exact jagged end length deduced by CC-tag strategy.











