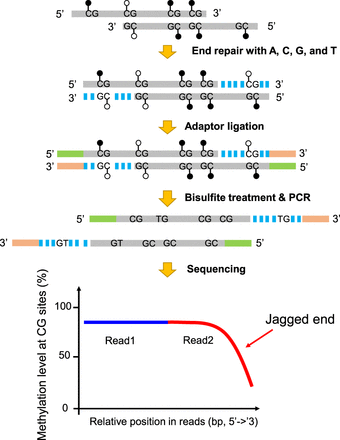
Schematic illustration of plasma DNA jagged end detection using methylation levels at CG sites. A DNA molecule carrying 5′ protruding ends (i.e., jagged ends) would be filled up by the unmethylated nucleotides (i.e., A, C, G, and T) with the presence of T4 DNA polymerase and the Klenow fragment, thus turning into blunt ends. The CG sites present in the newly generated strand were expected to be unmethylated, whereas the CG sites in the original sequence of the same molecule had, on average, a 70% chance of being methylated. The resultant blunted molecules were subjected to bisulfite treatment and PCR amplification. If a molecule contained jagged ends with a 5′ protruding single strand, the methylation levels at CG sites proximal to the 3′ end (e.g., read2) would be lower than that close to the 5′ end (e.g., read1). Filled lollipops represent methylated Cs, and unfilled lollipops represent unmethylated Cs. The dashed blue lines represent newly filled-up nucleotides.











