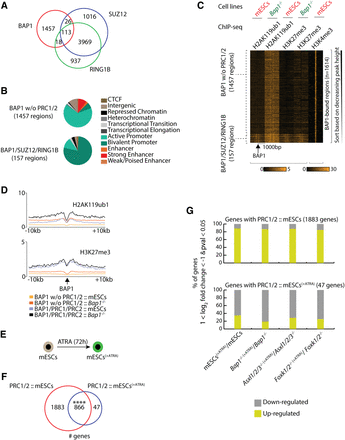
PR-DUB and PRC1/2 share a limited set of bivalent target genes. (A) Euler diagram showing the overlap between BAP1, SUZ12, and RING1B binding regions in wild-type proliferating mESCs. (B) Hidden Markov analysis for the 1457 BAP1-bound regions positions not associated with SUZ12 or RING1B in mESCs, and for the 157 BAP1/SUZ12/RING1B-bound regions in wild-type mESCs. The distribution of the regions is shown in the 11 indicated categories. (C) Heat maps showing the depicted ChIP-seq profiles (H2AK119ub1, H3K27me3, and H3K4me3) in 2 kb centered around the BAP1-bound regions in wild-type and Bap1−/− mESCs. (D) The average signal of H2AK119ub1 and the H3K27me3 binding profile shown in a window of 20 kb around the BAP1-bound regions with or without SUZ12/RING1B in wild-type and Bap1−/− mESCs. (E) Schematic representation of mESCs subjected to all-trans retinoic acid (ATRA)-induced differentiation for 72 h (mESCs[+ATRA]). (F) Euler diagram showing the overlap of PRC1/2 target genes, identified in proliferating mESCs and in mESCs treated with ATRA for 72 h (mESCs[+ATRA]). Fisher's exact test, (****) P-value < 0.0001. (G) Expression changes of genes in response to ATRA-induced differentiation in wild-type, Bap1−/−, Asxl1/2/3−/−, and Foxk1/2−/− mESCs. Upper panel shows the % of the 1883 genes that are bound by PRC1/2 only in mESCs and are up- or down-regulated in response to ATRA. Lower panel shows the expression changes of the 47 genes that are only bound by PRC1/2 in mESCs treated with ATRA for 72 h (mESCs[+ATRA]).











