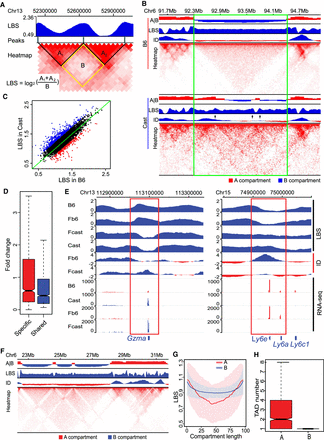
Strain-specific TAD boundaries and relationships between TAD and A|B compartment. Resolution: LBS (5 kb); ID (5 kb); Heatmap (20 kb); compartment (100 kb). (A) Definition and calculation of LBS. (B) LBSs could accurately detect the TAD boundaries. The green box outlines a region with several strain-specific TAD boundaries (black arrows) and divergent A|B compartments. (C) Significantly different TAD boundaries between B6 and Cast inferred by LBS. (D) Genes near strain-specific TAD boundaries showed stronger strain-specific expression than these near strain-shared TAD boundaries: (Specific) strain-specific TAD boundaries; (Shared) strain-shared TAD boundaries. (E) Strain-specific boundaries are associated with strain-specific gene expressions near Gzma and Ly6e, respectively. The red box highlights strain-specific boundary. (F) LBSs in B compartments are flat and smooth, whereas LBSs in A compartment fluctuate. (G) Comparison of mean and SD of LBSs in A and B compartments, showing a high variability of LBS on A compartments. (H) Box plot of number of TADs in A compartments and B compartments (P-value < 2.2 × 10−16).











