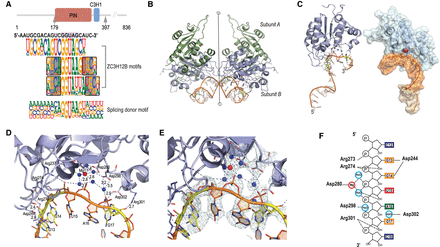
Structural basis of RNA motif recognition by ZC3H12B. (A) Schematic representation of the domain structure of ZC3H12B. The arrows indicate the first and the last amino acid of the construct used for crystallization, containing both the PIN domain (residues 181–350) (Senissar et al. 2017) and the known RNA-binding C3H1 zinc finger domain (residues 355–380) (Lai et al. 2002; Hudson et al. 2004). RNA sequence used for crystallization and all ZC3H12B motifs and for the splice donor motif are shown below the cartoon. Note that all these motifs contain the sequence GGUA. (B) Figure shows two asymmetric units of the crystals of RNA-bound ZC3H12B (a total of two RNAs, four ZC3H12B proteins). Only the PIN domain is visible in the structure. The crystals belong to the P43212 space group, and the asymmetric unit contains one protein dimer of two identical monomers presented in green (subunit A) and blue (subunit B) and one RNA molecule. This dimer is similar to the dimer found in the structure of ZC3H12A (PDB: 3V33) (Xu et al. 2012). Note that the contact between the two dimers of ZC3H12B around the twofold crystallographic axis (vertical line) is primarily mediated by the two RNA chains. Red and blue spheres represent Mg2+ ions and water molecules, respectively. For clarity, only the water molecules found in the active site are shown. Dashed lines represent hydrogen bonds (right side). The residues involved in the protein–RNA contacts are shown as ball-and-stick models, and the nucleotides involved in hydrogen bonds with these residues are in yellow. Notice that only the active site of subunit B of the AB dimer is occupied by an RNA molecule. (C) The structure of ZC3H12B PIN domain. (Left) The PIN domain is composed of a central beta-sheet surrounded by alpha-helices from both sides. The RNA molecule is bound near the Mg2+ ion by the -GGUAG- sequence, which is located close to the 3′ end of the cocrystallized RNA. (Right) Surface model shows the shape of the active site bound by RNA (brown), with the weakly coordinated Mg2+ ion. Waters are omitted for clarity. Note the horseshoe-like shape of the RNA backbone at the active site (orange). (D) A closeup image of the RNA fragment bound to the catalytic site of ZC3H12B. Mg2+ ion is shown as a red sphere; the water molecules are represented as blue spheres, with dashed lines representing hydrogen bonds. Note that phosphates of U15, A16, and G17 interact with the Mg2+ ion via water molecules. The Mg2+ ion is coordinated by five water molecules that also mediate contact with one of the side-chain oxygen atoms of Asp280 as well as Asp195 and Asp298 and phosphate groups of RNA. Thus, the octahedral coordination of the Mg2+ ion is distorted, and the ion is shifted from the protein molecule toward the RNA chain, interacting with the RNA via an extensive network of hydrogen bonds. The RNA backbone is slightly bent away from the protein, suggesting that the sequence is a relatively poor substrate. The presence of only one magnesium ion and the positions of water molecules correspond to the cleavage mechanism suggested for the HIV-1 RNase H (Keck et al. 1998). (E) The image in D annotated with the 2Fo−Fc electron density map contoured at 1.5 σ (light green mesh). (F) Schematic representation of interactions between protein, the Mg2+ ion, and RNA. Solid lines represent contacts with RNA bases, whereas hydrogen bonds to ribose and phosphates are shown as dashed lines. Nucleotide bases are presented as rectangles and colored as follows: G, yellow; A, green; U, red; and C, blue. Water molecules and Mg2+ ion are shown as light blue and red rings, respectively.











