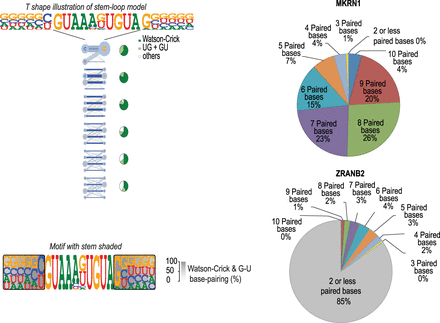
Comparison between linear PWM and stem–loop (SLM) models. (Left) Visualization of the stem–loop models. (Top) A T-shape model shows a horizontal loop and a vertical stem where the frequency of each base combination is shown. Bases are aligned so that Watson–Crick base pairs orient horizontally. Pie-charts show frequency of Watson–Crick (green) and G-U base pairs (light green) compared with other pairs (gray) that do not form canonical dsRNA base pairs at each position of the predicted stem. (Bottom) A linear visualization in which the base-pairing frequency is indicated by the darkness of gray shading is also shown. (Right) RNA secondary structure prediction analysis using RNAfold reveals that sequences flanking MKRN1 loop sequence form base pairs (top), whereas bases on the flanks of ZRANB2 matches (bottom) are mostly unpaired.











