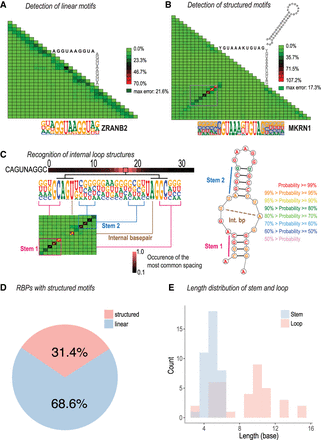
Detection of linear or structured RNA-binding models. (A) ZRANB2 binds to a linear RNA motif. The motif of ZRANB2 and the seed used to derive it are shown below and above the triangular correlation heat map, respectively. The heat map illustrates deviation of the observed nucleotide distributions from those predicted by a mononucleotide model in which bases are independent. (B) MKRN1 binds preferentially to a stem–loop. Note a diagonal series of red tiles (boxed) that indicates pairs of bases whose distribution deviates from the independence assumption. These bases are shaded in the motif below the triangle. The interdependency occurs between bases that are at the same distance from the center of the motif, consistent with formation of a stem–loop structure. Right top: An RNAfold-predicted stem–loop structure for a sequence that was highly enriched in the experiment. (C) LARP6 binds to a complex internal loop RNA structure. The left panel indicates the dinucleotide dependencies with the heat map on top representing the preferred spacing length between base-pairing sequences of stem 1, whereas the right panel presents a predicted structure of the bound RNA. The dashed line in the structure denotes the internal base pair. (D) Fraction of RBPs with linear and structured binding specificities. RBPs with at least one structured specificity are counted as structured. (E) Length distribution of stem and loop for the structured motifs.











