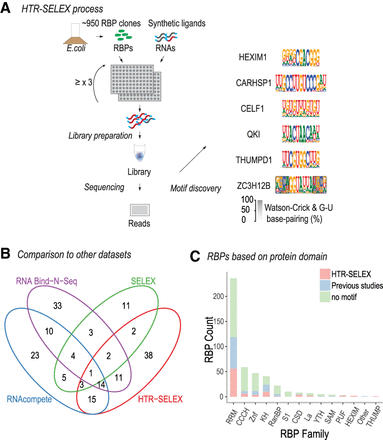
HT RNA-SELEX protocol and data analysis. (A) Schematic illustration of the HTR-SELEX process. RBDs or full-length RBPs expressed in E. coli as TRX-HIS6-SBP–tagged fusion proteins (top left) were purified and incubated with barcoded RNA selection ligands. RNA ligands bound by the proteins were recovered by RT-PCR, followed by in vitro transcription to generate the RNA for the next cycle of SELEX (middle left). The procedure was repeated at least three times, and the ligands recovered from the selection cycles were subjected to Illumina sequencing (bottom left) with data analysis to generate binding specificity models (right). (B) Comparison of the number of RBPs with motifs derived in the present study (HTR-SELEX) with the number of RBPs for which motifs were previously derived using RNA Bind-n-Seq (RBNS) (Dominguez et al. 2018), SELEX, and/or RNAcompete (cisBP-RNA version 0.6) (Ray et al. 2013). Note that our analysis revealed motifs for 38 RBPs for which a motif was not previously known. (C) Distribution of RBPs with motifs classified by the structural family of their RBDs. RBPs with motifs reported by Ray et al. (2013) and Dominguez et al. (2018) are shown in blue, and RBPs for which motifs were not reported there but determined using HTR-SELEX in this study are in red. RBPs with no motifs are in green.











