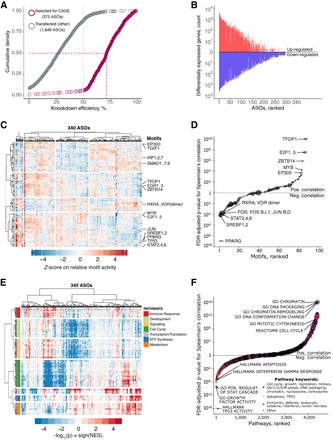
CAGE predicts cellular phenotypes. (A) RT-qPCR knockdown efficiency for 2021 ASO-transfected samples (targeted lncRNAs only). Gray dashed line indicates 50% KD efficiency generally required for CAGE selection. Purple dashed lines indicate median KD efficiency (71.5%) for 375 ASOs selected for CAGE sequencing. After quality control, 340 ASOs targeting lncRNAs were included for further analysis. (B) Distribution of significantly differentially expressed genes (up-regulated: FDR < 0.05, Z-score > 1.645, log2FC > 0.5; and down-regulated: FDR < 0.05, Z-score < −1.645, log2FC < −0.5) across all 340 ASOs. (C) Motif Response Activity Analysis (MARA) across 340 ASOs. Scale indicates Z-score of the relative motif activity (the range was set to abs[Z-score] = <5 for visualization purposes). (D) Correlation between normalized growth rate and motif activities across 340 ASOs targeting lncRNAs with highlighted examples. Motif sizes shown are scaled based on the HDF expression of their associated TFs (range 1 to ∼600 TPM). (E) Enriched biological pathways across 340 ASOs. Scale indicates GSEA enrichment value calculated as −log10(p) × sign(NES). (F) Same as in D but for selected GSEA pathways. Pathways sizes are scaled based on the number of associated genes.











