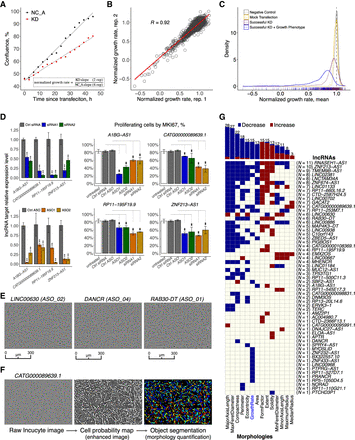
Cell growth and morphology assessment. (A) Selected example (PTPRG1-AS1) showing the normalized growth rate estimation using a matching NC_A (negative control). (B) Correlation of the normalized growth rate for technical duplicates across 2456 Incucyte samples. (C) Density distribution of normalized growth rates (technical replicates averaged) 252 ASOs targeting lncRNAs with successful knockdown (KD) and growth phenotype (blue) consistent in two replicates (FDR < 0.05 as compared to matching NC_A; 246 ASOs inhibited growth), 627 ASOs targeting lncRNAs with successful KD (purple), 270 negative control (NC_A) samples (gray), and 90 mock-transfected cells (Lipofectamine only) samples (yellow). (D) MKI67 staining (growth inhibition validation) for four selected lncRNA targets after siRNA and ASOs suppression. (E) Incucyte cell images of selected distinct cell morphologies changes upon an lncRNA KD. (F) An overview of the cell morphology imaging processing pipeline using a novel lncRNA target, CATG000089639.1, as an example. (G) lncRNAs (N = 59) significantly (FDR < 0.05) and consistently (after adjusting for the number of successfully targeting ASOs) affecting cell growth (N = 15) and cell morphologies (N = 44).











