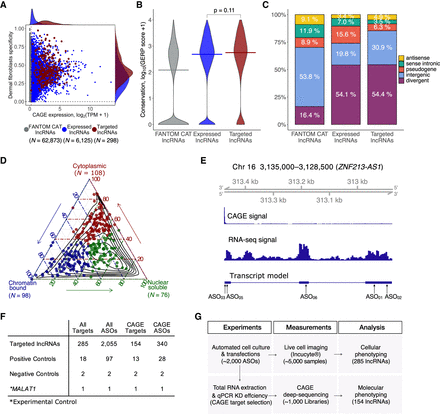
Selection of lncRNA targets, their properties, and the study overview. (A) CAGE expression levels at log2TPM (tags per million) and human dermal fibroblasts (HDFs) specificity of lncRNAs in the FANTOM CAT catalog (Hon et al. 2017) (N = 62,873; gray), lncRNAs expressed in HDFs (N = 6125; blue), and targeted lncRNAs (N = 285; red). The dashed vertical line indicates most lowly expressed lncRNA target (∼0.2 TPM). (B) Gene conservation levels of lncRNAs in the FANTOM CAT catalog (gray), lncRNAs expressed in HDFs (blue), and targeted lncRNAs (red). Crossbars indicate the median. No significant difference is observed when comparing targeted and expressed in HDF lncRNAs (Wilcoxon P = 0.11). (C) Similar to that in B but for genomic classes of lncRNAs. Most of the targeted lncRNAs and those expressed in HDFs are expressed from divergent promoters. (D) Subcellular localization (based on relative abundances from RNA-seq fractionation data) for targeted lncRNAs. Chromatin-bound (N = 98; blue); nuclear soluble (N = 76; green); cytoplasmic (N = 108; red). Black contours represent the distribution of all lncRNAs expressed in HDFs. (E) Example of ZNF213-AS1 loci showing transcript model, CAGE, and RNA-seq signal along with targeting ASOs. (F) Number of ASOs for target lncRNAs and controls used in the experiment. (G) Schematics of the study.











