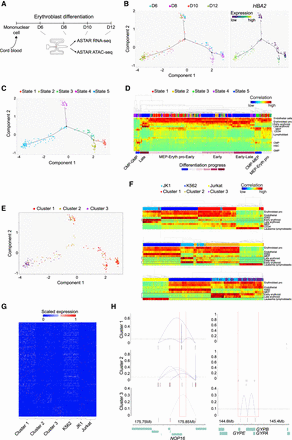
Interactive analysis of transcriptome and chromatin accessibility during erythroblast differentiation. (A) Schematic of erythroblast differentiation time points harvested for ASTAR-seq library preparation. (B, left) Trajectory of erythroblast differentiation identified from ASTAR RNA-seq libraries using DDRTree dimension reduction. Colors represent time points. (Right) Superimposition of HBA2 expression on the trajectory. Colors represent expression levels. (C) Trajectory plot revealing the pseudotemporal states. Colors represent pseudotemporal states. (D) RCA heatmap showing the clustering of cells undergoing erythroblast differentiation, based on their correlation to the cells of different lineage origins in RCA panel. Color indicates correlation value, ranging from blue (low) to red (high). Each row indicates one lineage, whereas each column represents an ASTAR RNA-seq library. Pseudotemporal state of each cell is indicated on top. Cellular differentiation status is determined based on their correlation to the cells of RCA panel and indicated below. (E) Superimposition of NMF clusters on the trajectory plot. (F) RCA clustering of JK1, K562, and Jurkat cells with the cells of NMF cluster 1 (top), cluster 2 (middle), and cluster 3 (below), respectively. Color indicates correlation value, ranging from blue (low) to red (high). Each row indicates one lineage, whereas each column represents an ASTAR RNA-seq library. Cell identities are indicated on top. (G) Heatmap showing the expression of NMF cluster–specific genes across NMF clusters and the hematopoietic cell lines. Color indicates expression level, ranging from blue (no) to red (high). (H) Line plots indicating the Cicero coaccessibility links between the regions highlighted in red and the distal sites in the surrounding regions. Height indicates the coaccessibility score of the connected peaks. The links are constructed from ASTAR ATAC-seq libraries of cluster 1 (top), cluster 2 (middle), and cluster 3 (bottom), respectively.











