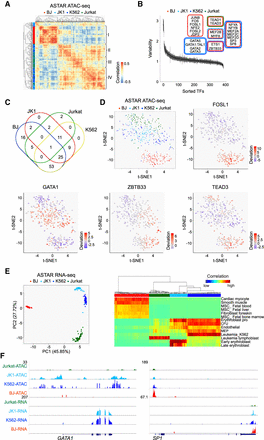
Application of ASTAR-seq on human cell lines. (A) Clustering of BJ, JK1, K562, and Jurkat ASTAR ATAC-seq libraries based on the human JASPAR motif deviation scores calculated over the HARs. Color indicates the correlation level among the libraries, ranging from blue (no) to red (high). Side color bar (y-axis) indicates the identity of each cell. (B) Variability plot indicating the variable TF motifs across the ASTAR ATAC-seq libraries of four human cell lines. The y-axis represents the variability score assigned to each JASPAR motif, whereas the x-axis represents the motif rank. Top variable motifs are classified based on their enrichment scores across the cell lines and colored accordingly. (C) Multi-Venn diagram showing the shared and unique TFs across the cell lines. (D, top left) t-SNE clustering of BJ, JK1, K562, and Jurkat ASTAR ATAC-seq libraries based on the deviation scores of human JASPAR motifs. Colors represent the cell lines. (Top right and bottom) Superimposition of motif enrichment scores for FOSL1, GATA1, ZBTB33, and TEAD3 on the t-SNE plot. Colors represent the motif enrichment levels, ranging from blue (no) to red (high). (E, left) PCA clustering of ASTAR RNA-seq libraries based on their correlation to the RCA panel. (Right) Heatmap showing the lineages that each cell correlates to. (F) UCSC screenshots indicating the chromatin accessibility (top) and expression (bottom) levels of GATA1 (left) and SP1 (right) across the cell lines.











