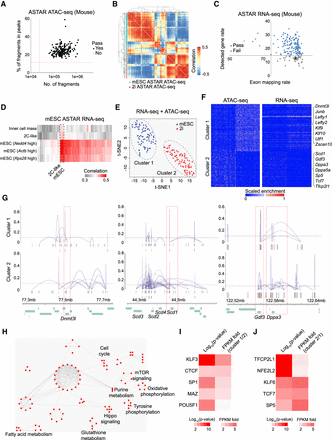
Transcriptomic and epigenetic heterogeneity within primed and naive mESCs. (A) Dotplot revealing the percentage of fragments in peaks (y-axis) and number of fragments (x-axis) of each mouse ASTAR ATAC-seq library. Red dotted lines represent the thresholds set for each criterion. (B) Heatmap showing the correlation among mESCs and 2i cells based on their ASTAR ATAC-seq libraries. (C) Dotplot revealing the detected gene rate (y-axis) of each mouse ASTAR RNA-seq library plotted against its exon mapping rate (x-axis). Blue dots represent the libraries that pass QC, whereas gray dots indicate the low-quality libraries. (D) Heatmap revealing the correlation of each mESC ASTAR RNA-seq library (x-axis) to various lineages of MCA panel (y-axis). Color indicates the correlation level, ranging from gray (low) to red (high). Two-cell-like (2C-like) mESCs are boxed with a dotted line. (E) NMF clustering of mESCs and 2i cells based on the correlative signals of their ASTAR ATAC-seq and ASTAR RNA-seq libraries. (F) Heatmaps revealing pairs of accessible regulatory regions (left) and the corresponding target genes (right) that are differentially enriched between the NMF clusters. Each column represents a library, whereas each row represents a chromatin region (left) or a gene (right). Color indicates the accessibility (left) and expression (right) levels, ranging from blue (low) to red (high). Representative genes are indicated on the right. (G) Line plots showing the differential coaccessibility links between the highlighted regions and its surrounding regions, identified using Cicero. Top plots are constructed from ASTAR ATAC-seq libraries of cluster 1 cells, whereas bottom plots are constructed from that of cluster 2 cells. Peak heights (y-axis) denote the coaccessibility scores. (H) Interactome analysis revealing the top pathways enriched by cluster 2–specific genes. (I,J) Heatmaps showing the enrichment (left columns) of TF motifs on cluster 1–specific (I) and cluster 2–specific (J) accessible regions and their relative expressions (right columns).











