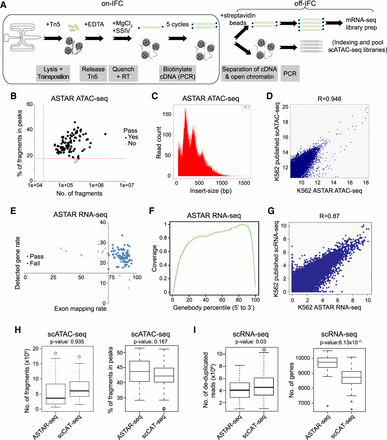
Assay for single-cell transcriptome and accessibility regions (ASTAR-seq). (A) Overview of ASTAR-seq protocol. (B) Dotplot revealing the percentage of fragments in peaks (y-axis) against the number of fragments (x-axis) of each K562 ASTAR ATAC-seq library. Red dotted lines represent the threshold values set for each criterion. (C) Histogram showing the frequency (y-axis) of fragments with the indicated insert size (x-axis). (D) Dotplot showing Pearson's correlation between K562 ASTAR ATAC-seq and the published K562 scATAC-seq libraries. (E) Dotplot revealing detected gene rate (y-axis) of each K562 ASTAR RNA-seq library plotted against its exon mapping rate (x-axis). Blue dots represent the libraries that pass QC, whereas gray dots represent the libraries of low quality. (F) Line plot representing the coverage ratio (y-axis) of K562 ASTAR RNA-seq reads over the gene bodies of housekeeping genes (x-axis). (G) Dotplot showing Pearson's correlation between K562 ASTAR RNA-seq and the published K562 scRNA-seq libraries. (H) Boxplots showing the number of fragments (left) and percentage of fragments in peaks (right) of scATAC-seq libraries prepared by the scCAT-seq and ASTAR-seq protocol. Two-tailed Student's t-test is used to calculate P-values. (I) Boxplots showing the number of de-duplicated reads (left) and genes (right) detected in scRNA-seq libraries prepared by the scCAT-seq and ASTAR-seq protocol. Two-tailed Student's t-test is used to calculate P-values.











