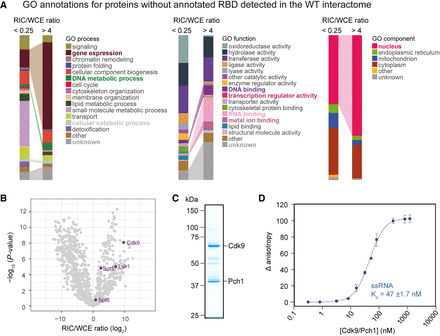
Characterization of RBPs with high RNA-binding activity that lack annotated RBDs. (A) Comparison of GO term distribution for RNA interactors without annotated RBDs that are either underrepresented (RIC/WCE ratio <0.25; n = 304) or enriched (RIC/WCE ratio >4; n = 203) in poly(A)+ RIC compared to statistical expectation (P < 0.01), visualized with the QuiLT tool on PomBase (Lock et al. 2019). Categories overrepresented among proteins enriched in RIC are highlighted in bold. (B) Volcano plot of the WCE-normalized RIC as in 1C. The cyclin-dependent kinases Cdk9 and Lsk1 and the transcription elongation factors Spt5 and Spt6 are enriched on poly(A)+ RNA. (C) Coomassie stain of Cdk9/Pch1 purified from insect cells. (D) Fluorescence anisotropy assay measuring binding of increasing amounts of purified Cdk9/Pch1 to 8 nM FAM-labeled RNA.











