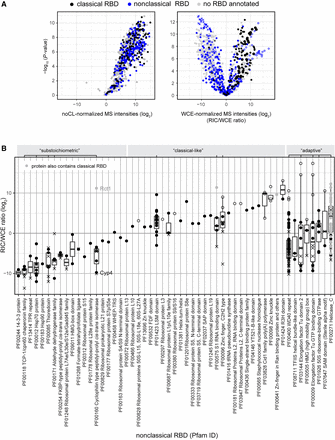
Properties of nonclassical RBDs. (A) Distribution of proteins annotated with a classical or nonclassical RBD in the noCL-normalized (left) and WCE-normalized poly(A)+ RNA interactome (right) as in Figure 1C. Nonclassical RBDs comprise 192 Pfam identifiers derived from the initial human interactome as well as any domain in RBDmap that had at least three peptides supporting it (Castello et al. 2012, 2016; Moore et al. 2018). The complete list of Pfam identifiers used for this analysis can be found in Supplemental Table S2. (B) Boxplot of RIC/WCE ratios for all proteins annotated with a nonclassical RBD that were detected in WT poly(A)+ RIC using symbols to indicate protein populations as in Figure 1D. Proteins that also harbor a classical RBD are marked in gray. Proteins annotated with GO:0022626 [cellular component: cytosolic ribosome] are not included. Abundant nonclassical domains (where at least four different domain-containing proteins were detected in the experiment) were classified as either “substoichiometric,” “classical-like,” or “adaptive,” as indicated above the plot.











