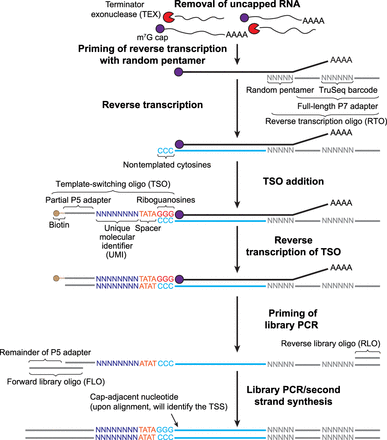
Schematic illustration of the STRIPE-seq method. Briefly, total RNA is treated with Terminator 5′-phosphate-dependent exonuclease (TEX) to reduce the proportion of uncapped RNA present in the sample. After a 1-h incubation, template-switching reverse transcription (TSRT) is performed using a barcoded reverse transcription oligo (RTO) primed with a random pentamer, followed by the addition of a unique molecular identifier (UMI)-containing, 5′-biotin-modified template-switching oligo (TSO) with three 3′ riboguanosines that permit the annealing of the oligo to the untemplated triplet Cs that are generated by reverse transcriptase when it reaches the m7G cap. Library PCR is then performed using the cleaned TSRT product as input, which ensures that TruSeq adapters are present on both sides of the insert (blue line). The cap-adjacent base, which identifies the transcription start site (TSS), is identified using the signature sequence [N8]-TATAGGG in the R1 read. Once the library PCR step is completed and cleaned, the STRIPE-seq library is then submitted for sequencing on an Illumina platform.











