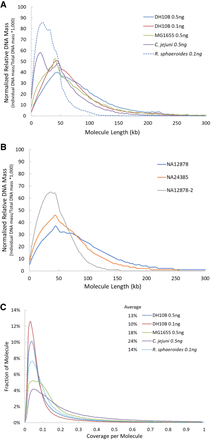
TELL-seq linked-read molecule analyses. (A) Calculated molecule length based on the TELL-seq sequencing data from microbial samples. To compare results from different microbial samples, the calculated DNA mass was normalized as follows: (Individual DNA mass at specified molecule length/Total DNA mass of the microbial sample) × 1000. (B) Calculated molecule length based on the TELL-seq sequencing data from human cell line samples. To compare results from different samples, the calculated DNA mass was normalized as follows: (Individual DNA mass at specified molecule length/Total DNA mass of the cell line sample) × 1000. (C) Distribution of linked-read sequencing coverage per molecule. Average sequencing coverage per molecule was 13%, 10%, 18%, 24%, and 14% for E. coli DH10B (0.5 ng genomic DNA input for library prep), E. coli DH10B (0.1 ng), E. coli K12 MG1655 (0.5 ng), C. jejuni (0.5 ng), and R. sphaeroides (0.1 ng), respectively.











