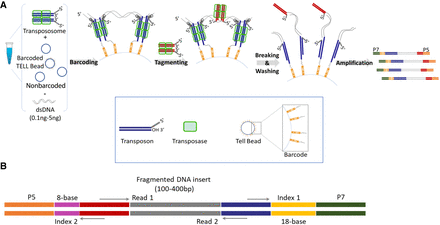
Overview of TELL-seq library workflow and structure. (A) Diagram of TELL-seq library preparation procedure. In a 0.2-mL PCR tube, 0.1 ng to 5 ng genomic DNA was mixed with 3–10 million barcoded TELL beads and transpososomes for the clonal barcoding reaction. Genomic DNA fragments were captured on the barcoded TELL beads via connecting strand transfer complexes (STCs) to barcode oligos on the bead surface. A tagging between STCs by a second transpososome introduced a second priming site for library amplification. After breaking the STCs and washing the magnetic TELL beads, sequencing library molecules were amplified off beads with P5 and P7 adaptor sequences incorporated at the same time. The total library procedure took ∼3 h. (B) TELL-seq library structure for Illumina sequencing systems. Index 1 comprises 18-base TELL-seq molecular barcode; Index 2 comprises 8-base barcode for sample indexing.











