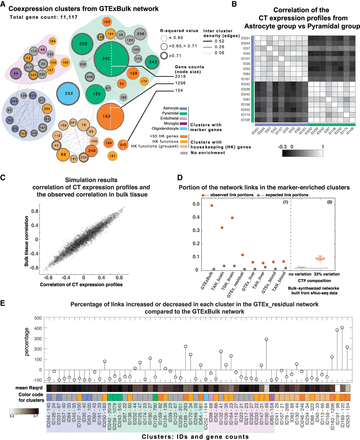
Much coexpression is explained by cellular composition effects. (A) Coexpression clusters from GTExBulk data set network. Clusters are labeled with their IDs. Color indicates if the cluster was enriched with markers of a specific cell type, is enriched with housekeeping functional terms, or has a high count of housekeeping genes. Thickness of the border reflects the mean R2 value for genes in the cluster. (B) Coexpression of mean CT expression profiles for a group of clusters affiliated with Pyramidal cell expression patterns and a group of clusters affiliated with Astrocyte cell expression patterns (affiliation is indicated with presence of markers, high R2, or high inter-cluster links). (C) Results from simulated bulk tissue data. Each dot represents data from a pair of genes. Plot shows data for 1000 gene pairs, sampled from a bulk tissue data set with 100 samples and 10 hypothetical cell types. As demonstrated, for a given pair of genes, their Pearson's correlation in the bulk tissue is highly correlated with the Pearson's correlation between their CT expression profiles. Also, the higher the correlation between their CT expression profiles, the more likely their correlation in the bulk tissue is the same as the correlation of their CT expression profiles. (D) Proportion of coexpression involving the set of genes from the clusters enriched with marker genes. Panel 1: The two brain networks (GTExBulk and the TAN-brain) and the brain-specific network (TSN-brain) have between ∼30% and 50% intra-cluster links in clusters enriched with marker genes. Panel 2: Portion of links in the same set of clusters in two groups of synthesized bulk tissue networks, modeling the effect of cellular composition variation. (E) Percentage of links increased or decreased for GTExBulk clusters in the residual network (GTEx_residual). Cluster color code as in A. Clusters are ordered based on their grouping in A. The range for average R2 is shown in the color bar. Many housekeeping clusters (orange) with low average R2 values yield more links in the residual network.











