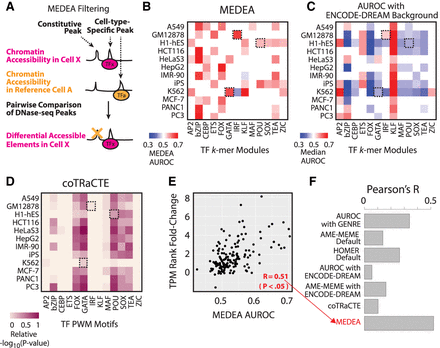
Motif enrichment of differential elements of accessibility (MEDEA). (A) Schematic for MEDEA peak filtering. Chromatin accessibility peaks of a cell type of interest (Cell X) and of a reference cell type (e.g., Cell A) are compared. Overlapping peaks, which indicate constitutively accessible regions, are subtracted out, thus only selecting the peaks specific to the cell type of interest (X not in A) that are likely bound by lineage-specific TFs (TFx). (B–D) For the same TF binding motifs and cell lines of Figure 1A, analysis of motif enrichment obtained by using (B) MEDEA (filtering and AUROC), (C) Glossary's AUROC using ENCODE-DREAM DNase-seq data sets as background sequences for enrichment calculation, and (D) coTRaCTE. Boxes highlighting known lineage specifiers as in Figure 1A. (E) For the TF families and cell lines used to benchmark MEDEA, scatter plots to correlate transcriptomic up-regulation (x-axis values from Fig. 1B) and MEDEA AUROC (Fig. 2B). (F) Barplot of the correlation coefficients (Pearson's R) between TF up-regulation and motif enrichment obtained from different methods (in red; R from MEDEA, as in E). For additional scatterplots depicting correlations for other methods, see also Supplemental Figure S6.











