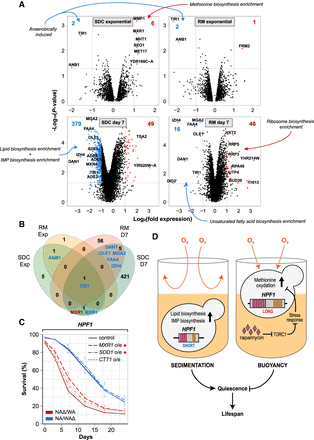
HPF1-induced buoyancy reprograms methionine, lipid, and purine metabolism. (A) Transcriptome changes induced by WA-HPF1-dependent buoyancy. NA/WA hybrids hemizygotes for WA or NA-HPF1 were cultivated in calorie-rich (left panels) or rapamycin (right panels) medium, and RNA was extracted and sequenced from exponential phase (top panels) or aging (bottom panels; 7 d after entry into quiescence) cells. y-axis: −log10 (P-value), x-axis: log2 (HPF1 NAΔ/WA) – log2 (HPF1 NA/WAΔ). Blue: transcripts more (>2×, P < 0.05) abundant in HPF1 NA/WAΔ, red: transcripts more (>2×, P < 0.05) abundant in HPF1 NAΔ/WA. The total number of transcripts passing each criterion are reported (top corners, blue and red text). Gene Ontology classifications enriched among transcripts passing each criterion are indicated (blue and red arrows) (Supplemental Table S5). (B) Comparing the number of transcripts more (>2×, P < 0.05) abundant in HPF1 NA/WAΔ (blue) or in HPF1 NAΔ/WA (red) across environments (SDC and RM, in exponential phase and after 7 d of aging [D7]). (C) CLS of HPF1 reciprocal hemizygotes overexpressing (o/e) either MXR1 (dashed), SOD1 (dot-dashed), or CTT1 (dotted) in SDC media in 96-well plate. Red asterisk indicates significance (P < 0.05 unpaired Student's t-test) at 7 d (MXR1) and 12 d (MXR1 and SOD1). (D) Model for how the intragenic tandem repeat expansions in WA-HPF1 shortens the life span by shifting cells from a sedentary (left) to a buoyant lifestyle (right), exposing them to more oxygen and causing mild oxidation of, in particular, methionine, which impairs entry into quiescence. Rapamycin prevents the accumulation of oxidized methionine, possibly through inhibiting TORC1 and through the activation of stress response genes.











