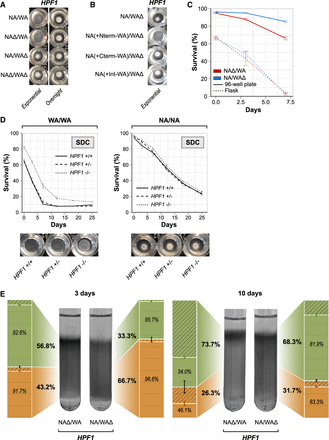
Buoyancy triggered by HPF1 N-terminal repeat expansions shortens life span. (A,B) Buoyancy of cells cultivated for 7 h (exponential phase) or overnight in calorie-rich medium in a 96-well plate. (A) HPF1 hemizygotes. (B) HPF1 allele swaps (as described in Fig. 3C). (C) Comparing CLS of HPF1 hemizygote cells cultivated in shake flasks and 96-well plates. Shake flasks had a 1:5 medium/volume ratio and were shaken at 220 rpm. Ninety-six-well plates were filled with 200 µL medium, with no shaking. (D) CLS and buoyancy (96-well plates; exponential phase) of WA and NA homozygotes parents with no (full line), 1 (dashed lines), or both copies (dotted lines) of HPF1 deleted in calorie-rich medium. (E) Percoll density gradients with HPF1 hemizygotes incubated in SDC media in a 96-well plate for either 3 d (left panel) or 10 d (right panel). The upper (nonquiescent cells) and lower (quiescent cells) phases were isolated by pipetting. The fraction (bold) and viability (italics) of cells in each phase were measured by flow cytometry (bar plots). Green: Upper/nonquiescent fraction, orange: lower/quiescent fraction, hatched area: dead cell fraction.











