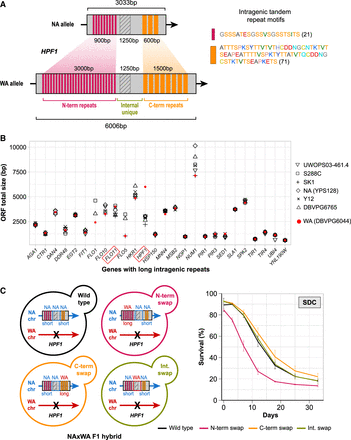
Massive intragenic tandem repeat expansions within HPF1 shorten life span. (A) Schematic representation of the intragenic repeats (colored rectangles) in HPF1 for NA and WA alleles. Hatched rectangle: a HPF1 internal unique region with high sequence variation between NA and WA alleles. Right: repeat motif units. Amino acids are colored according to the RasMol nomenclature. Numbers = motif size (amino acids). (B) Size variation of genes containing long intragenic repeats in seven diverged S. cerevisiae strains (Verstrepen et al. 2005; Yue et al. 2017). Diamonds: North American, red circles: West African. (C) Left panel: Design of allele swaps of HPF1 segments in the NA/WA F1 hybrid. The WA-HPF1 allele was deleted (black cross), while the NA-HPF1 was kept unchanged (wild type), or a segment was replaced by the corresponding WA-HPF1 segment. N-term: N-terminal repeats, C-term: C-terminal repeats, Int: internal unique region. Right panel: CLS for allele swapped constructs in SDC media.











