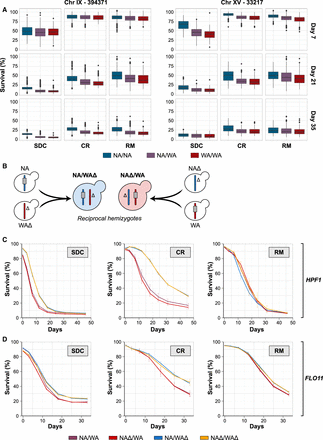
Natural allelic variations in the HPF1 and FLO11 control chronological life span. (A) Chronological life span of the 1056 POLs separated according to genotype at the markers with highest LOD score in each of the two major QTLs: 394,381 kb in Chromosome IX (in FLO11) and 33,217 kb in Chromosome XV (in HPF1). (B) Schematic representation of the NA/WA reciprocal hemizygosity design used to validate the CLS effect of the HPF1 and FLO11 WA alleles. Color: NA (blue) and WA (red) chromosomes. Gray rectangle: candidate gene (HPF1, FLO11), Δ: gene deletion. (C) Reciprocal hemizygosity. CLS of NA/WA hemizygotes for NA (blue; WAΔ) and WA (red; NAΔ) HPF1, heterozygote for HPF1 (purple; NA/WA), and lacking HPF1 (yellow; NAΔ/WAΔ). (D) As in C, but for FLO11.











