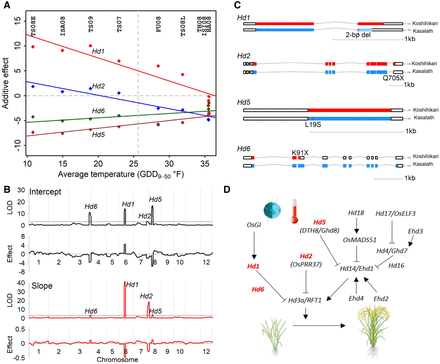
Varied effects at four loci along the temperature gradient underlying rice flowering-time plasticity. (A) Reaction norms of genetic effects at the single-locus level along the environmental index by temperature (GDD9–50). Dots show the effects detected from individual environment analysis. Dashed vertical line shows the positions of intercepts at the average temperature value. (B) Loci detected by mapping using the parameters (intercept and slope) derived from the reaction norms of genotypes. Horizontal lines in LOD plots show the significance threshold. Additive effect in descending absolute-value order for intercept (Hd5, Hd1, Hd6, and Hd2) and for slope (Hd1, Hd2, Hd5, and Hd6). (C) Known or potential functional polymorphisms between two alleles in four flowering-time genes from sequence analysis. (D) Positions of four genes highlighted in the rice flowering control pathway under natural long day-length (LD) conditions. These genes originally discovered for photoperiodic response are found to be involved in sensing temperature differences in LD. Pathway diagram is modified following the Figure 1 in the work by Matsubara et al. (2014).











