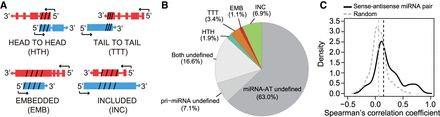
Global pattern of expression of sense–antisense miRNA pairs. (A) Schematic representation of different types of sense–antisense miRNA pairs. Depending on the positions of the transcripts involved, the pairs are classified as follows: (HTH) head-to-head (5′-regions overlap), (TTT) tail-to-tail (3′-regions overlap), (EMB) embedded (antisense transcript is fully contained within the sense transcript), and (INC) included (sense transcript is fully contained within the antisense transcript). The arrow indicates transcriptional direction. (Red) pri-miRNA; (blue) miRNA-AT; (slashed lines) pre-miRNA or the region opposite to the pre-miRNA. (B) The proportion of different types of pairs classified in A. (Undefined) no annotation was found for either pri-miRNA or miRNA-AT. (C) Distribution of Spearman's correlation coefficient between sense–antisense miRNA expression. A total of 130 sense–antisense miRNA pairs with measured expression levels in the FANTOM5 CAGE data were analyzed. Correlation between random pairs of miRNA and miRNA-AT genes is represented by a gray dashed line (n = 130).











