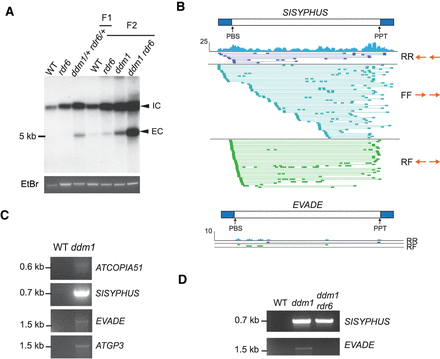
Extrachromosomal DNA of LTR retrotransposons in ddm1 and ddm1rdr6. (A) Southern blotting using an EVADE probe was performed with undigested genomic DNA of F1 and F2 plants from the same parental lines. Integrated DNA copies (IC) and extrachromosomal DNA copies (EC) are indicated. Ethidium bromide (EtBr) staining was used for loading control. (B) Discordant short-read alignments from SISYPHUS (AT3TE76225) and EVADE in ddm1. Read pair orientations (forward or reverse for the first and second mate): RR and FF reads align in the same direction to the reference, indicating inversions, whereas RF reads face outward, indicating circular templates. LTR regions are indicated as blue bars. (C) Inverse PCR with genomic DNA to detect circular extrachromosomal DNA from ATCOPIA51, SISYPHUS, EVADE, and ATGP3 in ddm1 plants. (D) Inverse PCR with VLP DNA and reverse-forward (RF) outward reading primers for SISYPHUS and EVADE. (C,D) PCR primers are listed in Supplemental Table S6.











