Figure 7.
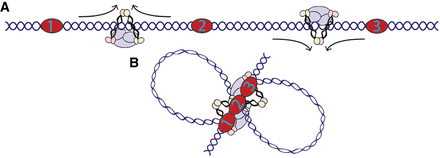
Model of condensin extrusion in C. elegans. (A) Extrusion in the X Chromosome of C. elegans likely begins at random locations and proceeds until blocked. Depicted here is bidirectional extrusion through one complex, but unidirectional extrusion through multiple complexes is also possible. (B) Loop anchors for DCC in C. elegans represent bidirectional blocks resulting in proximity of each anchor with every other.











