Figure 1.
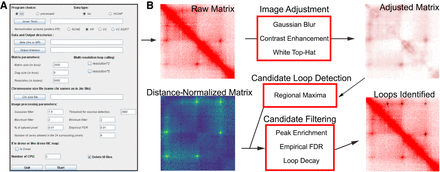
Overview of SIP. (A) The graphical user interface provides options to specify a Juicer-derived .hic file or processed data, the output directory, and chromosome size file. It also allows adjustment of the various parameters shown. (B) SIP uses image-based detection to create a long list of candidate loops, which is further filtered based on properties of the distance-normalized matrix.











