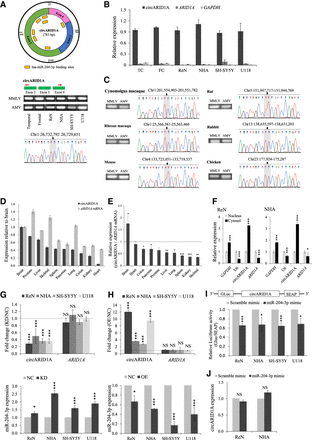
Experimental validation of the back-spliced junction of an up-regulated circRNA (circARID1A) and the corresponding circRNA-miRNA regulatory axis. (A) Validation of the NCL junction of circARID1A. (Top and middle) The schematic diagrams displaying seven predicted target sites of one single down-regulated miRNA (miR-204-3p) on the circle of circARID1A (top) and the designed divergent primers around the NCL junction of circARID1A (middle). (Bottom) Comparisons of two different RTase products (MMLV- and AMV-derived products) of circARID1A in TC/FC samples and four types of neuronal cell lines (hNPC [or ReN], NHA, SH-SY5Y, and U118), followed by Sanger sequencing the RT-PCR amplicons for the NCL event in the TC. (ReN) ReNcell VM. (B) Experimental validation of the circRNA (or back-spliced) junction of circARID1A. The figure shows the expression fold changes (as determined by qRT-PCR) for circARID1A, ARID1A mRNA, and GAPDH (negative control) in the indicated tissues/cell lines before and after RNase R treatment. (C) Experimental examination of the evolutionary conservation of circARID1A across the brains of vertebrate species from primates to chicken. Comparison of MMLV- and AMV-derived-RTase products (left) and the corresponding sequence chromatograms (right) for the circARID1A event in the brains of the indicated six species are shown. (D) Comparison of the expression profiles of circARID1A and its corresponding colinear mRNA counterpart in 10 normal human tissues. The expression levels of brain are used to normalize the relative expression values of the other tissues. (E) The relative expression of circARID1A and its corresponding colinear mRNA counterpart in 10 normal human tissues. (F) qRT-PCR analysis of the subcellular fractionation location for circARID1A and ARID1A mRNA. GAPDH and RNU6-1 snRNA are examined as a cytosol marker and a nucleus marker, respectively. The expression levels in nucleus are used to normalize the relative expression values in cytoplasm. (G,H) qRT-PCR analyses of the correlations between the expression of circARID1A and miR-204-3p after circARID1A knockdown (G) or overexpression (H) in various human neuronal cell lines. The top panels of G and H represent that circARID1A knockdown (G) or overexpression (H) did not significantly affect the ARID1A mRNA expression. (I) Luciferase reporter assay for the luciferase activity of GLuc-circARID1A in ReN, NHA, SH-SY5Y, and U118 cells transfected with miR-204-3p mimic and a scramble mimic (the negative control) to validate the binding between circARID1A and miR-204-3p. The entire circle sequence of circARID1A was cloned into the downstream region of the GLuc gene (i.e., GLuc-circARID1A; top). The luciferase activity of GLuc was normalized with secreted alkaline phosphatase (SEAP). (J) qRT-PCR analysis of the expression level of circARID1A in ReN and NHA cells after transfection with scramble and miR-204-3p mimics: (NC) negative control; (KD) knockdown; (OE) overexpression. All the qRT-PCR data are the means ±SD of three experiments. P-values are determined using two-tailed t-test. Significance: (*) P-value < 0.05; (***) P-value < 0.001; (NS) not significant.











