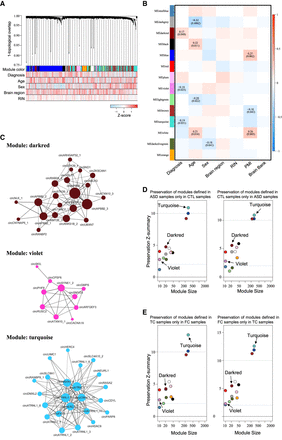
Dysregulation of circRNA coexpression networks in ASD cortex. (A) Dendrogram of circRNA coexpression modules defined in 134 cortex samples. Consensus module color bar shows assignment based on 1000 rounds of bootstrapping. Diagnosis and potential confounders (age, sex, region, RIN) are treated as numeric variables to calculate their Pearson correlation coefficients with expression level for each circRNA. (B) Pearson's correlation between distinct covariates and module eigengenes in 134 cortex samples. (C) circRNAs are plotted according to their circRNA expression correlations; the circRNAs in violet and darkred modules are all plotted, but the circRNAs in the turquoise module are plotted with only kME ≥ 0.5 due to the sizable number of circRNAs. Node size is proportional to connectivity, and edge thickness is proportional to the absolute correlation between two circRNAs. (D,E) Module preservation defined in ASD samples only in non-ASD samples (CTL samples) (D, left), CTL samples only in ASD samples (D, right), TC samples only in FC samples (E, left), or FC samples only in TC samples (E, right).











