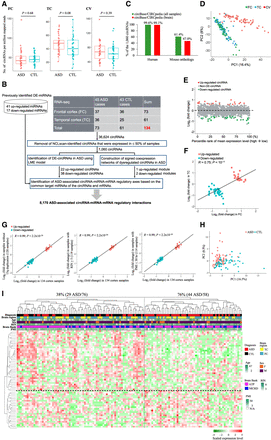
Identification of DE-circRNAs in the ASD cortex samples. (A) Comparison of normalized numbers of circRNAs in ASD and non-ASD control samples from different brain regions: (FC) frontal cortex, (TC) temporal cortex, (CV) cerebellar vermis. P-values were determined using two-tailed Wilcoxon rank-sum test. (B) Flowchart of the overall approach. To minimize potentially spurious events, only the circRNAs (1060 circRNAs) that were detected to be expressed in >50% of the samples examined are considered for the following analyses. (C) Comparisons of the 1060 circRNAs and the human/mouse circRNAs collected in well-known databases (i.e., circBase/CIRCpedia). (D) Principal component plots of circRNA expression profiles of the 1060 circRNAs in samples from FC, TC, and CV. (PC1/PC2) First and second principal components. (E) circRNA expression fold changes (>0 if higher in ASD; <0 if lower in ASD) between ASD and non-ASD control cortex samples, plotted against the percentile rank of mean expression levels of the 1060 circRNAs across 134 cortex samples used for differential expression analysis. The identified up-regulated (22) and down-regulated (38) circRNAs in the ASD cortex samples (LME model, P < 0.05 and |log2(fold change)| ≥ 0.5) are highlighted in red and green, respectively. (F) Comparison of circRNA expression fold changes in the FC and TC samples. The black line represents the regression line between fold changes in the FC and TC for the 60 DE-circRNAs. The Pearson correlation coefficient (R) and P-value are shown. (G) Comparison of circRNA expression fold changes in a small number of samples and all 134 samples combined: (left) samples with no Chromosome 15q11-13 duplication syndrome; (middle) samples with RIN ≥ 5; (right) samples with PMI ≤ 30 h). The black lines represent the regression lines between fold changes in the corresponding small number of samples and all samples combined for the 60 DE-circRNAs. (H) PCA based on the 60 DE-circRNAs. (I) Dendrogram representing hierarchical clustering of 134 cortex samples based on the 60 identified DE-circRNAs. Information on diagnosis, age, brain bank, PMI, brain region, sex, and RIN is indicated with color bars below the dendrogram according to the legend on the right. Heat map on the bottom represents scaled expression levels (color-coded according to the legend below the heat map) for the 60 DE-circRNAs.











