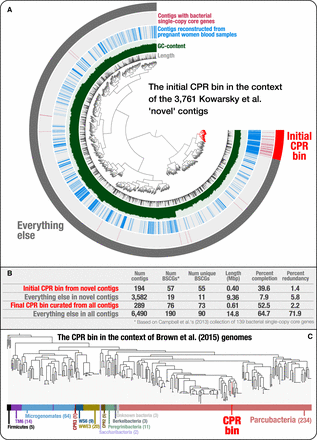
Genome-resolved metagenomics is essential to better investigate microbial diversity. (A) The inner dendrogram displays the hierarchical clustering of 3761 “novel” Kowarsky et al. contigs based on their tetranucleotide frequency (using Euclidean distance and Ward clustering) with the set of contigs that identify the genome in these data that is a member of the Candidate Phyla Radiation (CPR). Although the two inner layers display the length and GC content of each contig, the outermost layer marks each contig that contains one or more bacterial single-copy core genes. Finally, the second most outer layer marks each contig that originates from the assemblies of pregnant women blood samples. Although the pregnant women cohort was only one of four cohorts of individuals in Kowarsky et al. (2017) (others being heart transplant, lung transplant, and bone marrow transplant patients), most ribosomal proteins we found in the assembly originated from contigs that were assembled from the pregnant women (Supplemental Table S1). The signal in this layer shows that contigs with bacterial single-copy core genes associate very closely with other contigs based on tetranucleotide frequencies, and most of these contigs are assembled from pregnant women blood metagenomes, providing additional confidence that this group of contigs represents a single microbial population genome within the “novel” set of contigs that were released by Kowarsky et al. (2017) in their original publication. (B) Comparison of the initial CPR bin we have identified in the “novel” set of contigs to the final CPR bin we have refined using the entire set of contigs, which included non-novel contigs we obtained from the authors of the original study (M Kowarsky, J Camunas-Soler, M Kertesz, et al., pers. comm.). (C) Phylogenetic analyses show the placement of the CPR bin in the context of CPR genomes released by Brown et al. (2015). More details of this case study are available at http://merenlab.org/data/parcubacterium-in-hbcfdna/.











