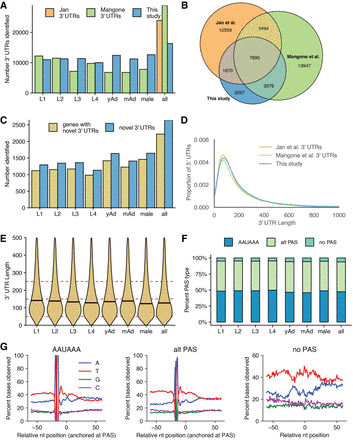
Properties of 3′ UTRome (A) Number of 3′ UTRs observed across all stages, as compared to Mangone et al. (2010) and Jan et al. (2011). (B) Venn diagram showing overlap between 3′ UTRs identified in this study, Jan et al., and Mangone et al. (C) Number of novel 3′ UTRs and genes with novel 3′ UTRs identified in each stage and across all stages. (D) Kernel density estimate plot of 3′-UTR lengths from this study, Jan et al., and Mangone et al. (E) Violin plots showing 3′-UTR length distributions across all stages. Horizontal black lines show the median of each stage. (F) Stacked bar chart showing percentage of UTRs with the specified polyadenylation signal (PAS) across all stages. (G) Nucleotide distributions around putative PAS sites and putative cleavage sites. Canonical PAS (AAUAAA) and alternative PAS (alt PAS) distributions are anchored with the putative PAS hexamer at −19 nt. The distribution of UTRs with no PAS is anchored with the putative cleavage site at 0.











