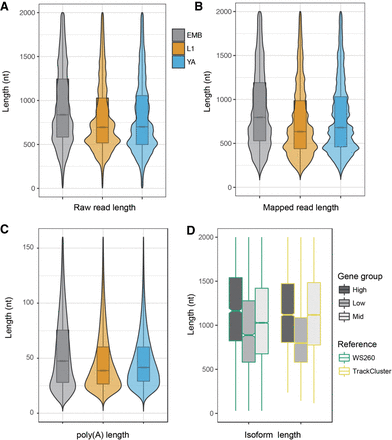
The long reads derived from embryos are significantly longer than those from L1 larvae or young adults. (A) Violin plots showing the length of raw reads derived from embryos (EMB), L1 larvae (L1), and young adults (YA). (B) Violin plots showing the length of mapped reads from EMB, L1, and YA stages. (C) Violin plots showing the length of poly(A) tails derived from EMB, L1, and YA stages. (D) Genes with a higher expression level (High) in embryo have an overall longer isoform. Box plots showing the length of isoforms from three categories of genes, that is, those that are expressed at a high (High), low (Low), or moderate (Mid) level in EMB compared to both L1 and YA stages (see Methods). Note that the genes with elevated expression in the embryo tend to have longer transcripts. The results from existing WormBase annotation (Release 260) and TrackCluster output are differentially color-coded in green and yellow boxes, respectively.











