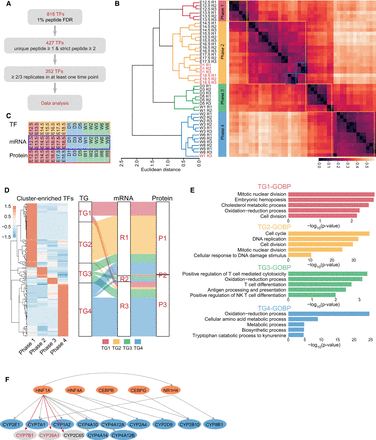
Analysis of transcription factors in developing mouse liver. (A) Flowchart for TFRE pulldown data quality control. (B) Hierarchical clustering analysis of the temporal expression of transcription factors separated the liver development into four phases. Each data point represents an independent replicate. (C) A comparison of developmental phases obtained by transcription factor binding activities, RNA expression, and protein expression. (D, left) Cluster-enriched phase-specific TFs; (right) a comparison of predicted downstream transcriptional targets (TG1–TG4) corresponding to TF Phases 1–4 and the distribution of these TGs detected in the RNA- or protein-based clustering. (E) Enrichment of the biological processes by the four TG groups. (F) Prediction of HNF1A as an upstream TF for CYP7B1 (protein and mRNA), CYP39A1 (protein and mRNA), and CYP2C65 (protein) by their coexpression with CYP family members known to be HNF1A TF targets (for heatmaps, see Supplemental Fig. S5D). (Orange) TFs; (blue) TGs known to be regulated by the TFs; (gray) TGs potentially to be regulated by the TF.











