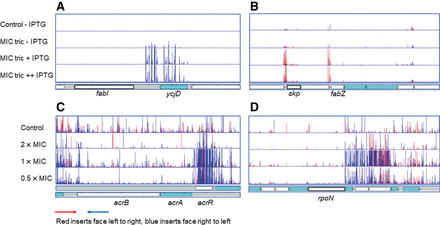
Validation of TraDIS-Xpress. Identification of known targets using the TraDIS-Xpress approach incorporating an inducible outward-facing promoter that identifies the impact of both essential and nonessential genes on survival and growth. (A) Genetic map of the relative gene positions is shown at the bottom of the panel. Above this, each row of vertical red or blue lines (plotted with red behind) indicates the position of mapped reads, and the height of the bar represents the relative number of reads mapped. Red indicates transposon insertions wherein the transposon-encoded kanamycin-resistance gene is oriented 5′ to 3′ left to right, and blue indicates the opposite right-to-left orientation. (A,B) Data split with induced and uninduced libraries shown at a single concentration of triclosan; this makes the impact of induction obvious. (C,D) Results from different drug concentrations (all with induction). (A) Inserts predicted to result in up-regulation of fabI in the presence of triclosan. (B) Mutants up-regulating skp, lpxD, fabZ, lpxA, and lpxB. (C) Mutants inactivating acrR and up-regulating acrAB being enriched by triclosan. (D) Mutants positioned antisense to rpoN being selected in the presence of triclosan. The top row in each plot shows untreated control cultures; the three rows below are for cultures grown in the presence of 2×, 1×, and 0.5× the MIC of triclosan (with IPTG induction for the outward facing promoter).











