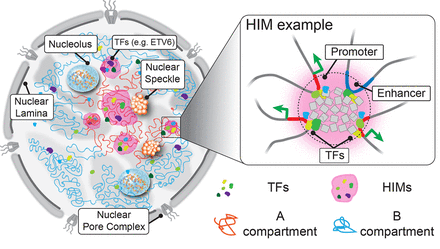
Illustration of the spatial organization of HIMs inside the nucleus. The cartoon on the left shows how chromosomes (curved lines) are intertwined in 3D space. Each chromosome can be primarily partitioned into an active A compartment (red) and an inactive B compartment (blue). Active and inactive genomic regions are formed in 3D space through cis- and trans-contacts, revealing shared localization relative to subnuclear structures, such as nuclear speckles and nuclear lamina. Similarly, the spatial localization of TFs within the nucleus is not randomly distributed but shows a great level of heterogeneity, probably affected by the distribution of binding sites on the 1D genome and the chromatin openness. As an example, ETV6 is highlighted. The MOCHI algorithm developed in this work is able to identify HIMs (shaded in pink), putative functional modules that transiently or stably exist in the nucleus, in which a group of TFs show an elevated concentration in a “transcriptional niche” and colocalize with genes in proximity in 3D. For example, a zoom-in view on the right reveals a potential scenario of a HIM in which the enhancer and its target genes located far away share binding by a group of TFs and are likely to be pulled together by TFs and cofactors. However, the exact interplay between TFs and 3D genome features and the global formation mechanisms of HIMs have yet to be revealed.











