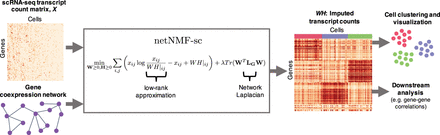
Overview of netNMF-sc. Inputs to netNMF-sc are: a transcript count matrix X from scRNA-seq and a gene coexpression network. netNMF-sc factors X into two lower-dimensional matrices, a gene matrix W and a cell matrix H, using the network to constrain the factorization. The product matrix 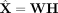 imputes dropped-out values in the transcript count matrix X. H is useful for clustering and visualizing cells in lower-dimensional space, whereas WH is useful for downstream analysis such as quantifying gene–gene correlations.
imputes dropped-out values in the transcript count matrix X. H is useful for clustering and visualizing cells in lower-dimensional space, whereas WH is useful for downstream analysis such as quantifying gene–gene correlations.











