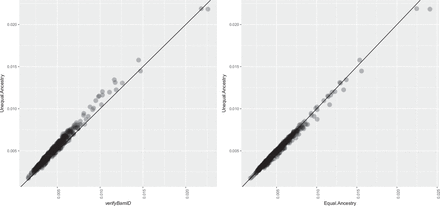
Comparison of contamination estimation between using verifyBamID and verifyBamID2 on 500 InPSYght samples. All subjects are African-Americans. Each dot represents the pair of contamination rate estimates using different methods. The left panel shows the estimated contamination rates of the original verifyBamID with pooled allele frequencies, which is the default setting of verifyBamID on the x-axis. The y-axis shows verifyBamID2 with unequal-ancestry model. Each point represents a sequenced subject. The right panel compares the estimated contamination rates between two models (unequal-ancestry vs. equal-ancestry) of verifyBamID2 on the same data set.











